After recording — compress, move, and unzip files:
- Zip and copy output files from Ultraspeech onto Z:/phon/data_upload/ (over the network)
- Using cox-515-1, open up the Y:\ (PHON) drive and run the archive extractor (this takes a few hours)
- Move the directory containing all the files into the directory where you will work with the data
- Also copy the log file from the /phon/may/Log_Files/ folder to the new working directory (so you don’t have to hunt it down):
cp /phon/may/Log_Files/LOG00733.TXT /phon/may/may33/
- After the extraction is complete, check to see if the the size of the wav file in bytes divided by 1470 is similar to the number of us images, and also to the number of video images, which you can check like this if you are in the /phon/may directory:
cd /phon/may ls may33/us_capture/ | grep -c bmp ls may33/video_capture/ | grep -c bmp ls -l may33/audio_capture
- If they don’t add up, alert Jeff.
Setting up and repairing the TextGrid:
- Run the
parselog.pyscript with “--beep 0” just to see the times when the beeps occur (after cd-ing into the participant’s main directory):cd /phon/may/may33 python /phon/scripts/parselog.py --input LOG00733.TXT --output may33 --beep 0 --carrier 'give me a X again'
- Running
parselog.pycreates a TextGrid that is not aligned. Open the TextGrid with the wav file and locate the last beep (which should be the occlusal plane in recent recordings) in the wav file, note its start time, then repeat the last step, replacing the 0 with the start time of the last beep. - (optional) offset: shift the prompt tier left (positive) or right (negative) by a fixed second duration in case there is a big alignment problem caused by speakers anticipating the prompts before they have pressed the forward button (which you will not find out until after the next couple steps).
python /phon/scripts/parselog.py --input LOG00733.TXT --output may33 --offset 0.0 --beep 42.4966 --carrier 'give me a X again'
- Generously segment the “palate” and “occlusal” intervals in the TextGrid phone tier, making note of the start and end times.
- Confirm that the palate and occlusal plane intervals are correctly aligned by using
makeclip.pyfor both, and checking out the movie files:python /phon/scripts/makeclip.py --wavpath audio_capture/may33.wav --start 9.9953 --end 14.1138 --output may33_palate python /phon/scripts/makeclip.py --wavpath audio_capture/may33.wav --start 41.4402 --end 44.375 --output may33_occlusal
- Once the beep is aligned, the prompts should be lined up under the correct trial. If this is not the case, check for a later beep (sometimes there’s accidentally a douple beep) and redo steps above. Once the prompts are lined up with each trial, use the forced aligner. Make sure you are still in the participant’s main directory, and run the following:
python /phon/p2fa/align_lab.py audio_capture/may33.wav may33_log.TextGrid may33.TextGrid
- First read may33.TextGrid in Praat, followed by may33_log.Textgrid, merge them, and save them as may33_merged.TextGrid in the participant’s main directory.
Correct the textgrid using the editor_align script
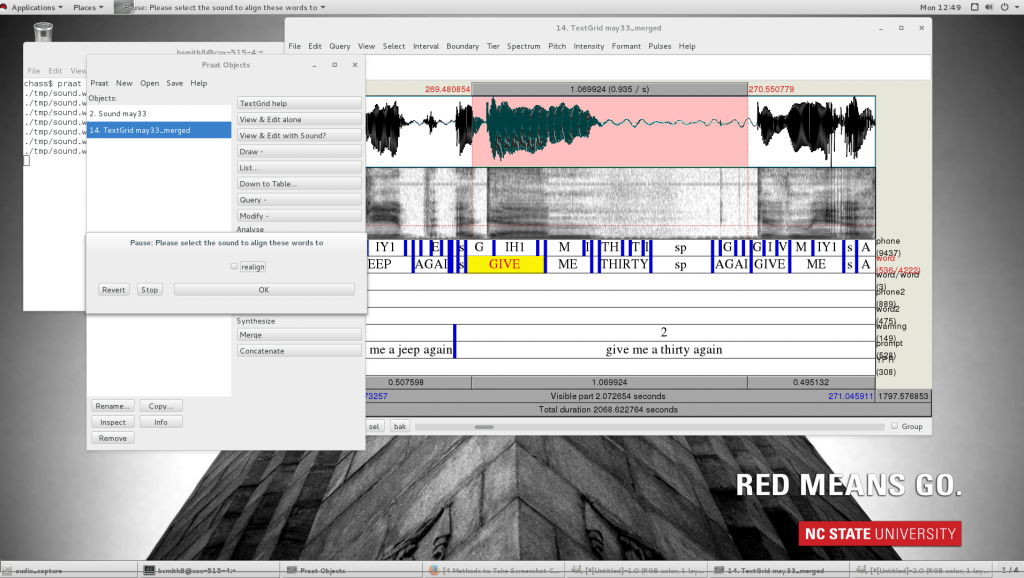
- Inside of the opened TextGrid editor, choose File>Open editor script… Select
editor_align.praatfrom the /phon/scripts directory. Then from inside the script window, choose File>Add to menu - Window should say TextGridEditor, the menu should be somewhere intuitive to look for it, for example, under Interval, and change the command to something you will recognize, such as “editor_align”
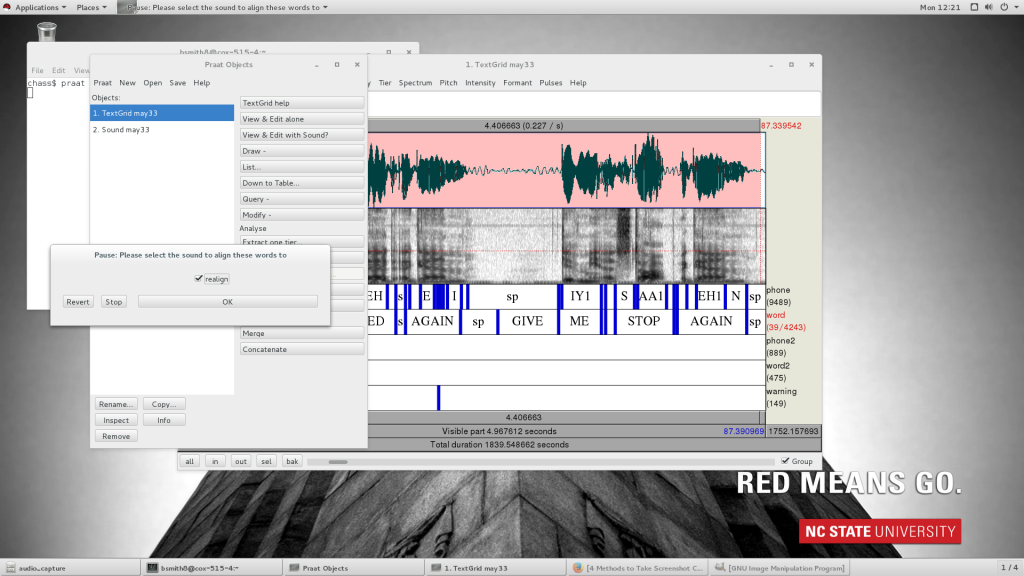
- Then when you see a misaligned section of phone tier, start by highlighting the interval that needs to be repaired, staying in tier 2. If you don’t have the correct section on the correct tier highlighted, it will delete things you don’t want deleted. Choose editor_align from the dropdown menu, and if you have excess word intervals where you don’t want them, uncheck the “realign” checkbox. This will remove all text from tier 1 and 2. (If it doesn’t, it’s because you’re in the wrong tier, so ctrl+Z to undo, then try again with the correct tier selected.)
- After removing excess words, you can replace any that you need to by simply typing it into an interval in tier 2, upper or lower case, entire phrases if you need to.
- Then choose Interval>editor_align again, after highlighting the phrases to be resegmented, and leave the checkbox ticked. This will fill in the phone tier from what you have selected in the word tier.
- Helpful hints for correcting TextGrids:
— Focus on prompts with warnings or prompts that overlap with the last part of a trial
— Remove target words that don’t belong in phone/word tiers (don’t worry about misaligned carrier phrases as long as they don’t interfere with the alignment of a target)
— Ignore words on phone2/word2 unless the important phones from the word or phrase are actually on-axis, and then you can selectively rehabilitate these by typing in the phrase in the appropriate interval on tier 2 and using editor_align.
— Make comments in warning tier if anything is really wierd, for example, if prompts don’t match the utterance that the participant produced.
— Save often. Praat likes to crash when you’re almost finished, so don’t let it take 3 hours’ work with it.
After the TextGrid has been corrected, start processing video and ultrasound
- The time stamps on each of the files are not all the same number of digits. To fix this, run
add_zeros.pyso that the naming conventions are all the same:python /phon/scripts/add_zeros.py us_capture/ python /phon/scripts/add_zeros.py video_capture/
- Use
bmpsmpl.pyto sample the video bmpspython /phon/scripts/bmpsmpl.py --output may33_video
- Open the resulting file, may33_video_summary.bmp, to help you choose a rectangular mask that includes all of the possible lip area, but cuts down on extra non-lip area.
- Each textured block in the summary output is 80×80 pixels, so add 80 pixels for each block, or 40 pixels for each half-block, etc. The four numbers you need to come up with are:
1) w= the width and 2) h= height of the box that will enclose the lips, and 3) the x- and 4) the y- coordinates for the top-left corner of the box, like this:python /phon/scripts/makepng.py --textgrid may33_merged.TextGrid --crop 360x280+160+80
Which will yield a box that is w=360 x h=280 pixels in size, starting at the coordinates x=160 and y=80.
makepng.pywill create two scripts that do the actual conversion: a test script, calledmay33_test(to see if you like your lip image cropping) and a convert scriptmay33_convert(to convert all the images for the palate and occlusal plane movies and the ultrasound and video, I think, maybe?).- Then you can check to see if you’ve cropped too much or too little by running:
./may33_test
and then look at images in /video_png to make sure they are the right dimensions and not cutting off any lip space. Re-run
makepng.pywith different dimensions, if necessary (you will have to delete the old files before you re-runmakepng.pybecause it will not overwrite existing files), and when you’re satisfied, run the following to convert all the rest of the files to pngs (using nohup so you can shut down your session while it runs):nohup ./may33_convert &
- To check whether the convert script is done (wait a few hours at least – imagine it’s like a 5-8 hour road trip, and you don’t want to start asking “Are we there yet?” until you’re at least halfway there), use the following command to find out: “how many images am I trying to convert?”:
grep -c png may33_convert
And then: “how many images have I created in the four directories that might hold images?”:
ls palate/ | grep -c png ls occlusal/ | grep -c png ls us_png/ | grep -c png ls video_png/ | grep -c png
Keep pasting these commands until the numbers add up to the number you got above. There should be a similar large number of us and video files, and a small number (a few hundred at most) of palate and occlusal files. When the numbers add up, it means all the images that were supposed to be created have been created. If you don’t have intervals in your textgrid’s phone tier labeled “palate” or “occlusal”, these directories will remain empty, so double-check the TextGrid and add them if this is the case. Run the script again if you have to, and don’t worry about re-writing the us and video files because the script won’t overwrite them. I think.
Create lists of relevant intervals to process
- Create folders to hold PCA output for us and video inside of the participant’s main directory:
mkdir us_pca mkdir video_pca
- Use
list_files_for_pca.pyto make filelists for ultrasound and video, which will be a list of which intervals to use and which to ignore. For the ultrasound, you don’t need to specify the--imagePathand--pca_pathbecause the default is to use the us_png and us_pca. You do need to add these arguments for the video.python /phon/scripts/list_files_for_pca.py --textgrid may33_merged.TextGrid --basename may33 --carrier 'GIVE ME A;AGAIN;GIVE;ME' python /phon/scripts/list_files_for_pca.py --textgrid may33_merged.TextGrid --basename may33 --carrier 'GIVE ME A;AGAIN;GIVE;ME' --imagepath video_png --pcapath video_pca
Then start preparing the ultrasound images
- Type this to enable MATLAB for this session:
add matlab81
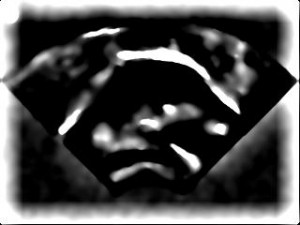
- The MATLAB functions will filter all of your images to enhance the tongue contour. The first step is to try two different filter settings and choose the best one. Enter this command, replacing all the instances of ‘may33’ to make the sample filtered images.
matlab -nodisplay -nosplash -nodesktop -r "P=path;path(P,'/phon/PCA');us_filter_check('may33','bmp','/phon/may/may33/us_png/','/phon/may/may33/us_pca/');exit" - This will produce mydata_LoG_1.jpg and mydata_LoG_2.jpg in /phon/may/may33/us_pca/ Download them and look at them, and decide which does the best job of enhancing the tongue surface without enhancing too much else. For example, see the sample filtered images below where the second (right-hand or lower) image is preferable because there’s not too much extra bright stuff that is not the tongue surface.
- Next, make a composite image from a sample of all your images. If you liked mydata_LoG_2.jpg the most, enter the following command with the 2 after ‘bmp’. If you liked mydata_LoG_1.jpg the most, put a 1 there instead.
matlab -nodisplay -nosplash -nodesktop -r "P=path;path(P,'/phon/PCA');us_selection_low('may33','png',2,'/phon/may/may33/us_png/','/phon/may/may33/us_pca/');exit" - Running the
us_selection_lowfunction will produce a file called may33_selection.jpg that is an average of a sample of your images, so that you can determine the range of tongue movements, and define a region of interest to restrict your analysis (so you don’t waste time calculating the values of pixels where the tongue never goes). To define the region of interest, first copy the selection file to the /phon/upload/ folder, renaming it test_selection.jpg:cp /phon/may/may33/us_pca/may33_selection.jpg /phon/upload/test_selection.jpg
- Choose the palate image by browsing to /phon/may/may33/palate and looking at the images, then copy it to the /phon/upload/ folder, renaming it test_palate.png.
cp /phon/may/may33/palate/may33_0010257_0587.png /phon/upload/test_palate.pngmay33_mask.csv
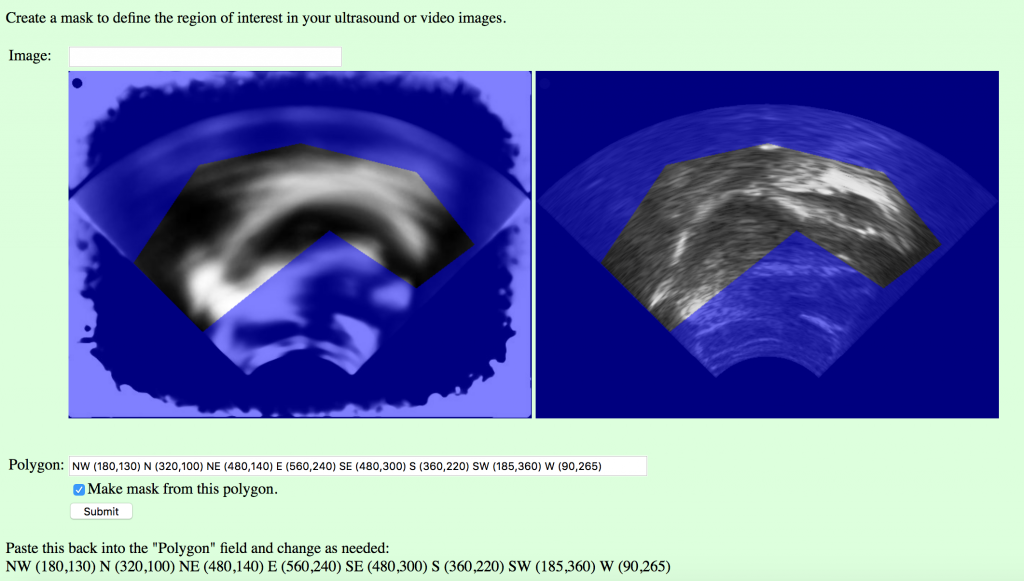
- Then go to the image mask page [http://phon.chass.ncsu.edu/cgi-bin/imagemask2.cgi] in a web browser. Enter “test_selection.jpg” in the “Image” field, and click Submit. You may need to refresh the page before the correct image appears. The two images will appear with a blue overlay (the mask), with the average tongue range on the left, and the palate frame on the right. The points that are written at the very bottom of the page (scroll down, if you have to), define the polygonal window in the mask (the region of interest) in a vector of (x,y) pixel coordinates. Copy the numbers into the “Polygon” field, make a change, and click “Submit” to see the result. You will have to keep pasting them back into the field to change them. The letters “NW”, etc. are just there to help orient you using compas points for up, down, left, and right, and you can use more or fewer points than the default of eight.
- When you have adjusted the points so that you can see the tongue through the window and the blue mask only covers the places the tongue doesn’t go, re-paste the coordinates and click the “Make mask from this polygon” checkbox and click submit one more time. (This would be a good time to paste the coordinates into the participants file, so you don’t have to look them up later.) Now there should be a test_selection_mask.csv file in the /phon/upload directory. This file will tell the next script which pixels to keep in each image. Copy it to your folder:
cp /phon/upload/test_selection_mask.csv /phon/may/may33/us_pca/may33_mask.csv
- For the lip video mask, MATLAB just needs to know the size and shape of the video clips that you chose earlier. So, rename the dummy mask file (which is just a list of the rectangular coordinates that got produced in the /us_pca folder when you ran the list selection script) to participantID_mask.csv:
mv /phon/may/may33/video_pca/dummy_mask.csv /phon/may/may33/video_pca/may33_mask.csv
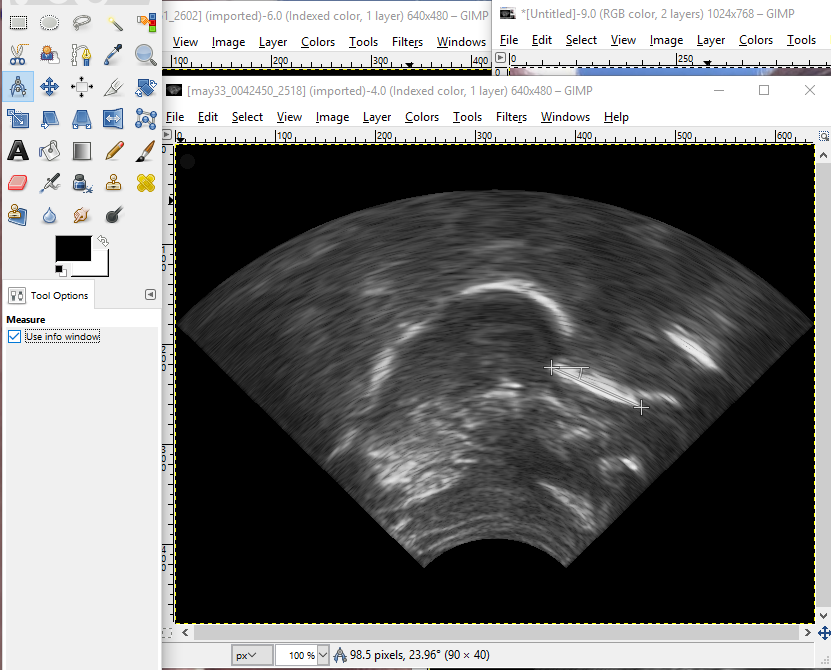
- One more thing you need to do before the PCA is to go back to the occlusal plane images in /phon/may/may33/occlusal and select one where there is a clear flat spot of the tongue against the tongue depressor. Open the file in GIMP, and use the measurement tool, as shown in the screen capture, to measure the angle of counter-clockwise rotation. Select the measure tool from the main toolbox or from the drop-down tools meanu. Make sure that “Use info window” is selected in the lower left area. Hold down the shift button while you click and drag the cursor across the flat edge of the tongue, and the number of pixels and angle (difference from 180 degrees) will be shown in the lower margin of the window. Here it shows 23.96, which we’ll round to 24 degrees.
Now we’re ready to do the Principal Components Analysis!
- There is a MATLAB pca function each for ultrasound and lip video. Both pca functions require a downsampling rate (both 0.2 below). This number determines how much the image resolution will be reduced. For this argument, use a value between 0 (no resolution) and 1 (full resolution). They also take an argument for the maximum number of principle components to calculate (both 50 below). If an integer value is used, the PCA function will retain that many PCs in the output. If, however, a non-integer value between 0 and 1 is used, the PCA function will interpret this number as a percentage threshold. In this case, the PCA function will retain as many PCs as are required to explain the given percentage of variance. For example, if you put 0.8 as a value for this argument, the output will retain as many PCs as are needed to explain at least 80% of the total variance in the image set.
- The ultrasound requires more information — 1) the counter-clockwise angle of occlusal plane rotation (24, in the example below, taken from the measurement in GIMP just above); use this argument to rotate images to the speaker’s occlusal plane. And filter (that was the contrast level (LoG) you chose earlier) as the third number after ‘png’.
- We can process about 10 frames per second (but we record at 60 frames per second), so processing takes about six times real time. That’s we run MATLAB using the command “nohup” (no hangup), so that the process will keep running even if you lose your connection to the server. These two commands create a script with all the arguments and files that you need to be able to call up using nohup so they can run
video_pca.mandus_pca.min the background:echo "P=path;path(P,'/phon/PCA');video_pca('may33','png',0.2,50,'/phon/may/may33/video_png/','/phon/may/may33/video_pca/');exit" > /phon/may/may33/video_pca/pca_cmd echo "P=path;path(P,'/phon/PCA');us_pca('may33','png',2,0.2,24,50,'/phon/may/may33/us_png/','/phon/may/may33/us_pca/');exit" > /phon/may/may33/us_pca/pca_cmd - Then you can execute them using
nohup. The following lines just execute the scripts you created in the above steps. Log onto a different server to run each of them to expedite the process (keep track of which machine each is running on so you can track errors).video_pcatakes about a half hour, whileus_pcashould take somewhere under 5 hours:nohup matlab -nodisplay -nosplash -nodesktop < /phon/may/may33/video_pca/pca_cmd & nohup matlab -nodisplay -nosplash -nodesktop < /phon/may/may33/us_pca/pca_cmd &
Merge video (you can do this at any time after the png files are created, but now is a good time)
- You can do this before or while the pca is running, since it’s not dependent on it. First, copy the merge_video_settings.txt file from may29 to the new participant’s folder:
cd /phon/may/may33 sed 's/may29/may33/' </phon/may/may29/merge_video_settings.txt >/phon/may/may33/merge_video_settings.txt
- Then merge the video:
nohup python /phon/scripts/merge_video_dev.py &
- After completion, find out how many files are in the us_with_video folder, and put that number in the “list” column in the Google spreadsheet:
ls us_with_video | grep -c jpg
Clean-up: make sure all the files are executable by the rest of the group
- If you are the owner of the directory, this is easiest for you to fix!
Make the directory executable (=usable as directories)chmod g+x /phon/may/may33
Make the subdirectories (and files) executable too (because there isn’t a good way to single out just the directories)
chmod g+x /phon/may/may33/*
Make the files with extensions non-executable (subdirectories and may33_test and may33_convert remain executable)
chmod g-x /phon/may/may33/*.*
Make the directory and all its contents readable (Recursively)
nohup chmod -R g+r /phon/may/may33 & (There will also be instances where we want directories and files to be writable by the group, but it's safest not to do that by default).
Make RData files for that participant
- Once everything is finished processing, and all the files are there and sorted out, open an R session in your terminal window, and collect the PCA data to attach to the RData file for that participant:
R setwd('/phon/may/RData') library(bmp) library(tidyr) # requires stringi library(tuneR) library(textgridR) # library(devtools); install_github("jpellman/textgridR") library(MASS) source('/phon/scripts/image_pca_functions.r') subjects <- c('may33') for (subject in subjects){ all_pca <- gather_pca(subject, perform_pca=TRUE) save(all_pca, file=paste0(subject,'.RData')) all_pca[[subject]]$us_info$vecs_variable <- NULL all_pca[[subject]]$video_info$vecs_variable <- NULL save(all_pca, file=paste0(subject,'_no_vecs.RData')) }
That’s it! It only took a few days, but you’ll get faster next time 🙂
List of scripts required for may protocol:
/phon/scripts/merge_video_dev.py
/phon/PCA/video_pca.m
/phon/PCA/us_pca.m
/phon/PCA/us_selection_low.m
/phon/PCA/us_filter_check.m
/phon/scripts/list_files_for_pca.py
/phon/scripts/bmpsmpl.py
/phon/scripts/makepng.py
/phon/scripts/makeclip.py
/phon/scripts/editor_align.praat
/phon/scripts/parselog.py
/phon/pf2a/align_lab.py
/phon/scripts/add_zeros.py
/phon/scripts/image_pca_functions.r